Research Vision of the Kastritis Laboratory for Biomolecular Research
Enzymopathies are metabolic disorders, often genetic, resulting from missing or defective enzymes. In order to prevent, combat or repair defective enzymes that lead to disease, it is vital that we understand the structure, function and interactions of enzymes and their complexes, ideally within their native cellular context.
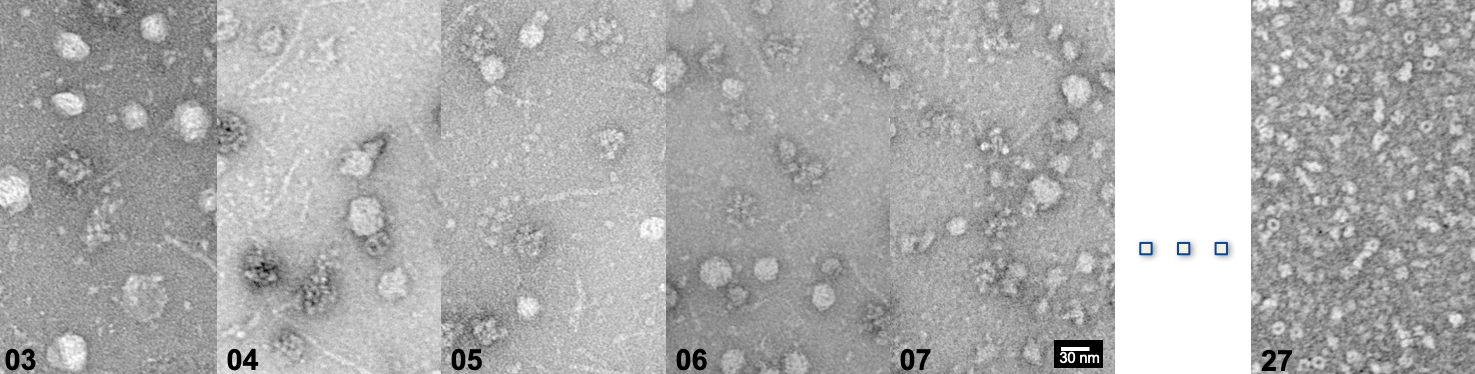
In their native cellular environment, enzymes are neither isolated nor randomly distributed. Instead, they co-compartmentalize, and often transiently interact to form supercomplexes of metabolic pathways (commonly referred to as metabolons). The formation of metabolons allows the intermediate product from one enzyme to be passed directly into the active site of the next consecutive enzyme of the metabolic pathway. Their role and existence were biochemically known for years but due to their sheer size and transient nature, understanding their molecular organization remains a challenge. As a consequence, traditional structural biology methods have helped us in understanding the structure and function of their constituent enzymes but not their higher-order assembly and how their interfaces are formed.
My research vision is to understand the molecular architecture of metabolons by integrating structural, biophysical, biochemical and biocomputational methodologies. This multidisciplinary approach aims at elucidating the molecular mechanisms that underpin cellular respiration in atomic details. In the long term, these methodologies will allow discerning cellular metabolons by visualizing their components and examining the interactions between the molecules that form the assemblies.
This research will ultimately provide new crucial insights not only to cellular respiration but also an entirely new view of cell mechanisms in general. The unsolved mystery of temporal and spatial synchronization of a myriad of protein subunits to perform a specific cellular function may be rationalized by the formation of supercomplexes. These unprecedented entities provide a whole new avenue for targeted drug design. Rather than targeting the active sites of individual subunits -which share many physical and chemical characteristics thus compromising specificity- I envisage a therapeutic strategy that targets the interface between proteins in the supercomplex that are likely to be unique.