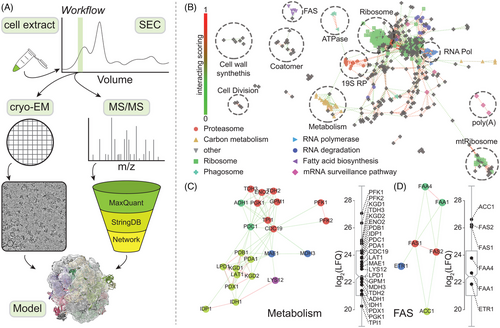
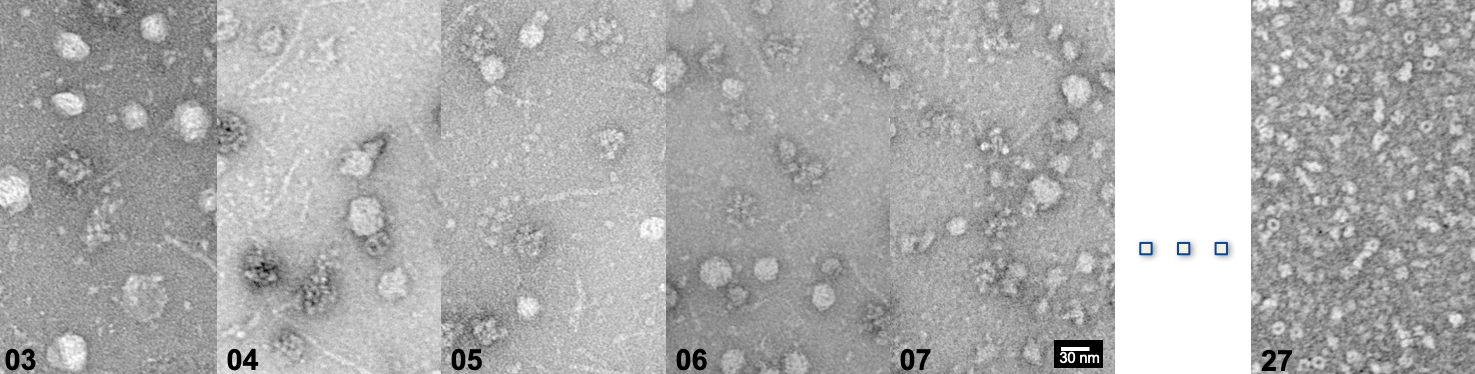
In a collaborative effort with the Sinz lab, our Post-doc Christian led a project on the utilization of AI-derived protein modeling based on mass spectrometry data, in order to interpret unknown cryo-EM maps from within native cell extracts!
This type of data integration is definitely the way to improve our understanding of native protein complexes. You can read all about this work HERE.
Very helpful guide! Many people still find it confusing to check their bantuan status. I also used this method and found it useful. For a quick check, I tried Cek Bansos and it worked well.
Download the latest version of Lucky PK Game for Android to access smooth gameplay, secure transactions, and exciting daily rewards.
Download the latest version of DK222for Android to access smooth gameplay, secure transactions, and exciting daily rewards.