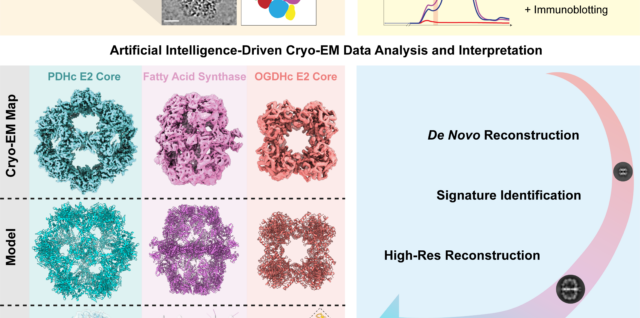
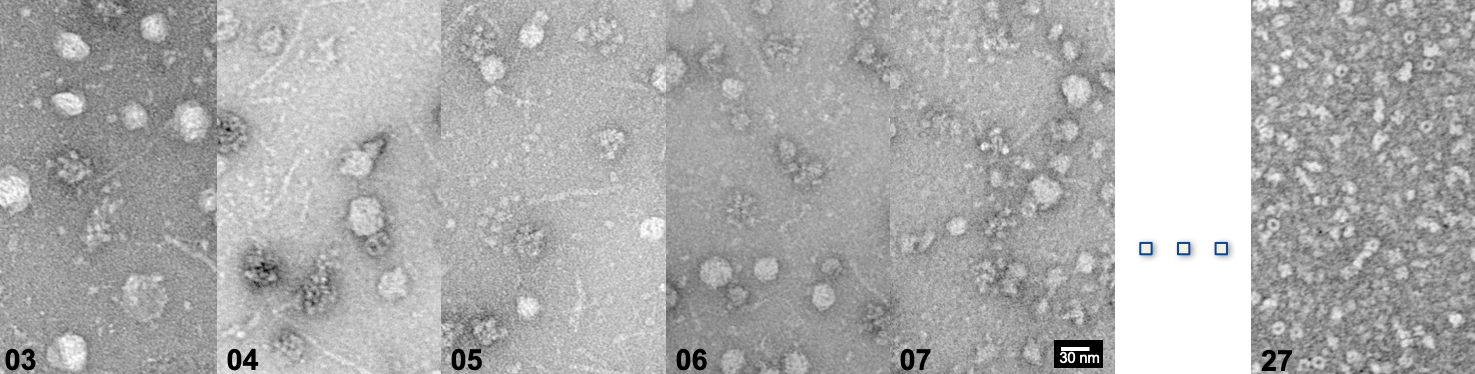
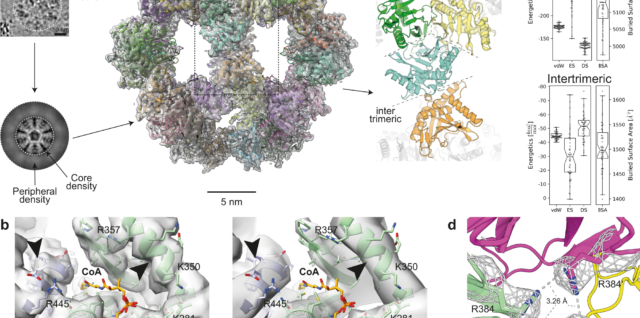
AlphaFold2 and Proteomics data help interpret native cell extracts
In a collaborative effort with the Sinz lab, our Post-doc Christian led a project on the utilization of AI-derived protein modeling based on mass spectrometry data, in order to interpret unknown cryo-EM maps from within native cell extracts! This type Continue reading AlphaFold2 and Proteomics data help interpret native cell extracts